Aims
The field of psychiatry is in dire need for a disorder classification system (a.k.a. diagnostic nosology) that is anchored in biological variables so that precise diagnosis, proper management of the disorder and efficient treatments could be developed. This study will challenge the existing group-based and descriptive approaches to analyzing neuroimaging data, by developing a computational approach to model how an individual’s brain- states change over time and to provide mechanistic guidelines as to how underlying neuronal processes attain such transitions (or lack thereof). The proposed computational psychiatric framework could not only be used for biologically grounded stratification of mental illnesses, but also as a test-bed for developing future treatments and person-centric care.
Here, we propose to take a bold step towards modeling the entire landscape of brain’s dynamical organization at an individual level. This modeled landscape could then be used as a “lens” towards (a) characterizing (and stratifying) psychiatric illness and (b) generating biologically grounded mechanistic insights regarding how neural processes interact during ongoing cognition to give rise to different dynamical landscapes in patient populations. We discuss several challenges associated with our approach, propose solutions, and provide initial proof-of-concept results. To test the efficacy of our approach in stratifying clinical populations, we aim to examine and stratify already collected data from neuro-dynamically opposite populations – Major Depressive Disorder (MDD) and Attention Deficiency and Hyperactivity Disorder (ADHD). Altogether, in this high-risk high-impact proposal, we challenge several existing paradigms to develop a computational psychiatry framework that will provide a much-needed platform for stratifying psychiatric illnesses and developing novel treatments to provide person-centric diagnosis and care.
Highlights
- Developed methods to characterize transitions in brain activity at the single participant level, without collapsing data in space or time at the outset
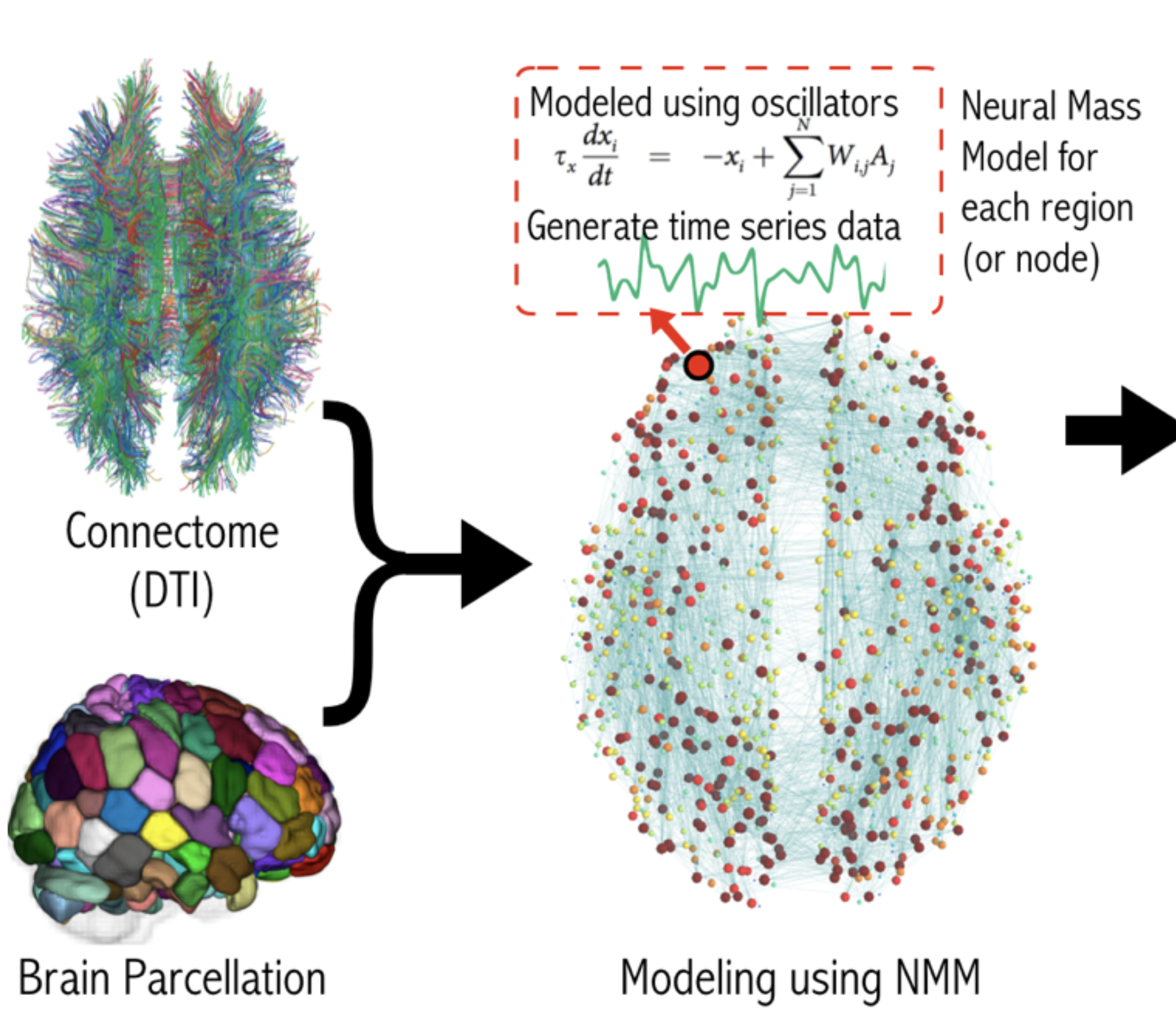
- Developed a new biophysically realistic model to better capture multiscale brain dynamics
- Developed new Topological Data Analysis (TDA) based methods to better capture transitions in brain dynamics
Presentations/Papers
- Maile, K., Saggar, M., R Miikkulainen. (2019) Implementing evolutionary optimization to model neural functional connectivity . Proceedings of the Genetic and Evolutionary Computation Conference Companion. ACM. 1731-33.
- Billings, J., Saggar, M., Hlinka, J, Keilholz, S., Petri, G. (2021). Simplicial and Topological Descriptions of Human Brain Dynamics. Network Neuroscience
- Saggar, M. & Uddin, L. (2019). Pushing the boundaries of psychiatric neuroimaging to ground diagnosis in biology . eNeuro.
- Geniesse, C., Sporns, O., Petri, G., Saggar, M. (2019). Generating dynamical neuroimaging spatiotemporal representations (DyNeuSR) using topological data analysis . Network Neuroscience.
- Zhang, M., Saggar, M. (2020). Complexity of resting brain dynamics shaped by multiscale structural constraints . BioRxiv.
- Saggar, M., Shine, J. M., Liégeois, R., Raut, R.V., Laumann, T.O., Snyder, A.Z., Dosenbach, N.U.F., Fair, D. (in-prep) Precision dynamical mapping using topological data analysis reveals a unique (hub-like) transition state at rest in highly sampled individuals.
- Giovanni, P., Saggar, M. (in-prep). Quantified persistence homology: stability and good practices for topological data analysis.
- Zhang, M., Chowdhury, S., Saggar, M. (in-prep). The topology of time: transition networks in simulated and real neural dynamics.
- Geniesse, C., Chowdhury, S., Saggar, M. (in-prep). Exploring the landscape of brain dynamics during ongoing cognition using topological data analysis and optimal transport.
Next steps
- Application of developmed methods to large-scale consortium data from clinical populations
Funding
This work was supported by the NIH Director’s New Innovator Award and MCHRI Faculty Scholar Fellowship to M.S.